Karolina came to us from Polish Academy of Science in Warsaw, Poland as a part of Fulbright’s BioLAB program and will stay with us for a year! Karolina will spend her internship on structural characterization of RNA-antibody complexes to gain better understanding on what structural features antibodies adopt to promote effective RNA binding. Welcome!
Author Archives: dankrochmal
New paper! | Structural Basis for Fluorescence Activation by Pepper RNA
Structural Basis for Fluorescence Activation by Pepper RNA
Pepper is a fluorogenic RNA aptamer tag that binds to a variety of benzylidene-cyanophenyl (HBC) derivatives with tight affinity and activates their fluorescence. To investigate how Pepper RNA folds to create a binding site for HBC, we used antibody-assisted crystallography to determine the structures of Pepper bound to HBC530 and HBC599 to 2.3 and 2.7 Å resolutions, respectively. The structural data show that Pepper folds into an elongated structure and organizes nucleotides within an internal bulge to create the ligand binding site, assisted by an out-of-plane platform created by tertiary interactions with an adjacent bulge. As predicted from a lack of K+ dependence, Pepper does not use a G-quadruplex to form a binding pocket for HBC. Instead, Pepper uses a unique base-quadruple·base-triple stack to sandwich the ligand with a U·G wobble pair. Site-bound Mg2+ ions support ligand binding structurally and energetically. This research provides insight into the structural features that allow the Pepper aptamer to bind HBC and show how Pepper’s function may expand to allow the in vivo detection of other small molecules and metals.

Allison Brink awarded Danute Nitecki Graduate Fellowship
Huge congratulations to Allison who just received Danute Nitecki Graduate Fellowship! Wishing you more successes on your PhD journey!
Congratulations to Dr. Huw Rees!
Huw just defended his PhD thesis! Congrats on your outstanding work on the crystal structure of Pepper RNA aptamer. We’ll miss you greatly and wish you all the best going forward!
New paper! | Structural basis for substrate binding and catalysis by a self-alkylating ribozyme
In January 2022 our group published a new paper in Nature Chemical Biology, where we describe a first crystal structure of a self-alkylating ribozyme along with some insights on the mechanism of reaction.
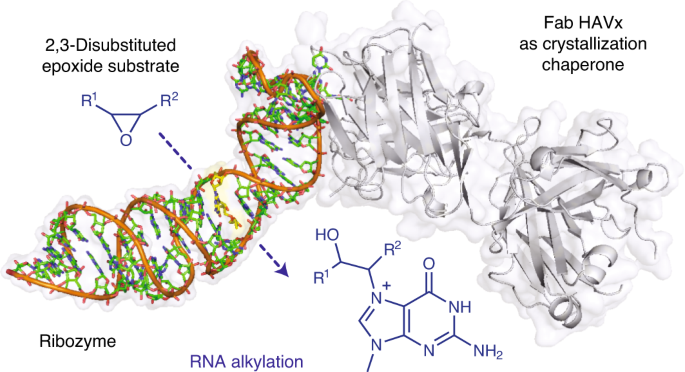
Structural basis for substrate binding and catalysis by a self-alkylating ribozyme
Ribozymes that react with small-molecule probes have important applications in transcriptomics and chemical biology, such as RNA labeling and imaging. Understanding the structural basis for these RNA-modifying reactions will enable the development of better tools for studying RNA. Nevertheless, high-resolution structures and underlying catalytic mechanisms for members of this ribozyme class remain elusive. Here, we focus on a self-alkylating ribozyme that catalyzes nitrogen–carbon bond formation between a specific guanine and a 2,3-disubstituted epoxide substrate and report the crystal structures of a self-alkylating ribozyme, including both alkylated and apo forms, at 1.71-Å and 2.49-Å resolution, respectively. The ribozyme assumes an elongated hairpin-like architecture preorganized to accommodate the epoxide substrate in a hook-shaped conformation. Observed reactivity of substrate analogs together with an inverse, log-linear pH dependence of the reaction rate suggests a requirement for epoxide protonation, possibly assisted by the ether oxygens within the substrate.
Christina Roman defended her PhD, congratulations!
Christina just defended her PhD thesis on her work on the structure of Programmed −1 Ribosomal Frameshifting Element from SARS-CoV2! Congratulations and all the best for you going forward!
Dan Krochmal awarded BSD Dean’s International Student Fellowship
Congratulations to Dan Krochmal, who has been awarded the BSD Dean’s International Student Fellowship!
Dan Krochmal joined the lab
Say hi to Dan Krochmal – our new PhD student! Dan Krochmal graduated with master’s degree from Jagiellonian University in Poland and visited the Piccirilli Lab in years 2017-2018 as a laureate of the Polish-American BioLAB program. Dan will be working on design and development of RNA-tailored phage-displayed Fab libraries.
Allison Brink joined the lab!
We’re happy to have a new PhD student in our ranks – say hi to Allison Brink! Allison graduated from Grinnell College and she will be working on atomic structure of Hovlinc ribozyme. Welcome!
Wojciech Gogacz joined our lab!
Piccirilli Lab is welcoming a new student Wojciech Gogacz, a chemistry graduate student at the University of Gdansk in Poland who is visiting us as a laureate of the Polish-American BioLAB program. Wojciech will spend one year in our lab working on structural investigation of Pepper fluorogenic RNA aptamer. Welcome!
